Malaria-Carrying Parasites Spread More When They Can Jump Into Multiple Birds - Study
 By Frank Otto
By Frank Otto

- ‘Too Much Going On’: Autistic Adults Overwhelmed by Nonverbal Social Cues
- Drexel’s College of Nursing and Health Professions Receives $1 Million for Scholarships from the Bedford Falls Foundation – DAF to Address Nursing Workforce Shortage
- U.S. Department of Education Provides Final Approval of Drexel and Salus Merger
- Spotting Bad Batteries Before They Malfunction
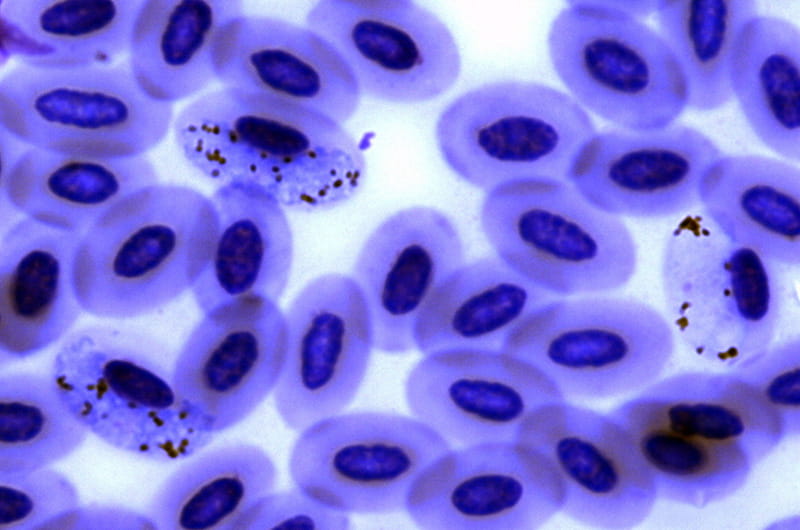
If you’re a parasite and want to spread out a little in Amazonia, then you better be cool with riding around in a variety of different birds, a new study found.
The Amazon region of South America is one of the most diverse tropical forest ecosystems on Earth, but an Academy of Natural Sciences of Drexel University study showed that in the blood parasites that cause malaria, bird hosts and the region’s physical geography play significant roles in the spread and diversity of these parasites.
A team led by Alan Fecchio, a former postdoctoral researcher at the Academy, and Jason Weckstein, PhD,an associate curator of the Ornithology at the Academy and an associate professor in Drexel’s College of Arts and Sciences, looked into two groups of haemosporidian parasites: Plasmodium and Haemoproteus.
“Plasmodium infection and distribution patterns are principally shaped by geography, whereas Haemoproteus patterns of diversity and distribution are primarily shaped by their host evolutionary relationships,” Weckstein said of the findings from his study, published in Ecography.
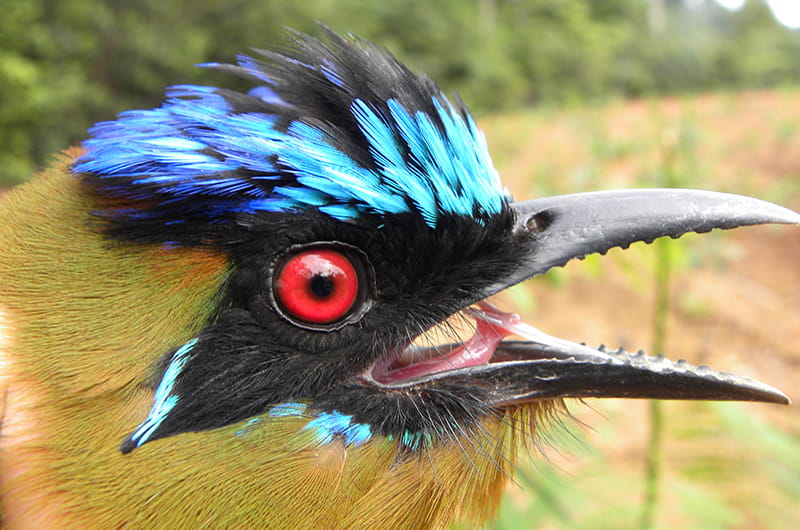

They found that the ability to switch to new bird host species was the prime indicator of regional diversity in haemosporidian parasites. Furthermore, Fecchio and colleagues found that Plasmodium, in particular, were more widespread, which makes sense because theyare “generalists” that can latch onto several different types of birds.
Haemoproteus, however, are limited in the types of birds they can infect. As such, they don’t get spread around very much.
But if Plasmodium can hop into a bunch of different birds, what keeps them from being everywherein this region? Birds can fly anywhere, right?
Well, it’s not as simple as that.

“Early 19th century Amazonian explorers Henry Walter Bates and Alfred Russel Wallace noted that major Amazonian rivers often appeared to form barriers to dispersal for birds, primates and other organisms,” Weckstein said. “As a result, if one compares the faunas found in areas flanking most Amazonian rivers, a large portion of the species will be different on opposite banks.”
Although the Amazon River is the most recognizable to most, there are many rivers in the Amazon region — such as the Xingu, the Tapajós and the Madeira. These make up roughly eight recognized areas of endemism in the region.
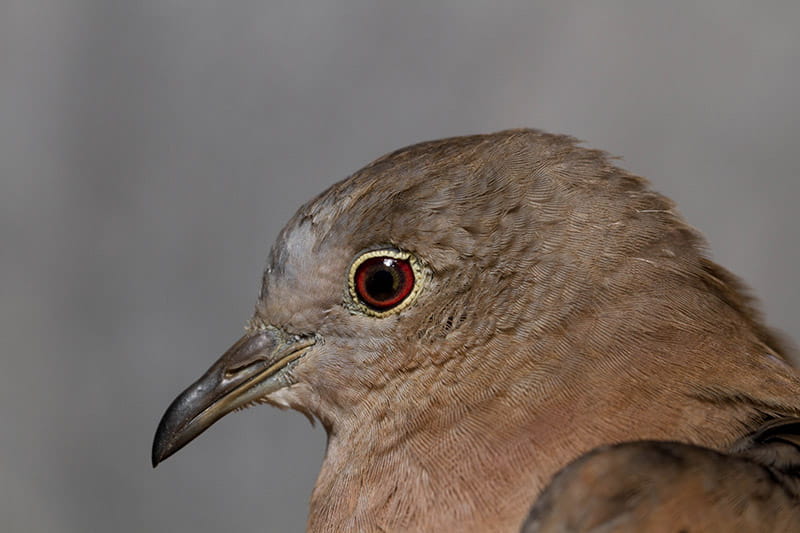
And, as it turns out, some of these birds just don’t have the endurance to make the trip across these rivers. Others might just have no reason to.
“Some birds never leave the dark understory of the forest and, thus, will not even cross small rivers, streams or open pastures,” Weckstein said. “Scientists have even conducted experiments and found that these understory birds fatigue and are unable to cross large expanses of open space, such as open water.”
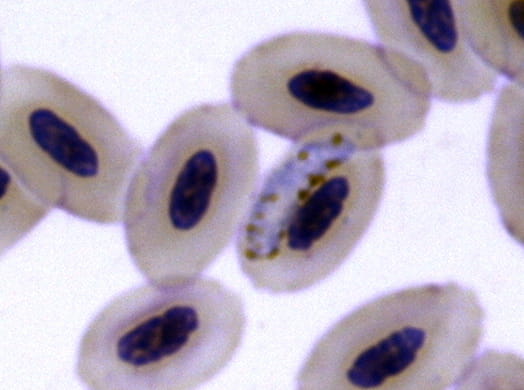
For comparison, some rivers in Amazonia are 10 to 20 times wider than the Delaware River at the spot where the Benjamin Franklin Bridge spans it in Philadelphia.
So even though Plasmodiumcan jump into multiple kinds of birds, if those birds are all staying put, so is the Plasmodium.
Although the study focused mainly on bird parasites, it provides another building block to understanding microorganisms that affect humans, too, like malaria. After all, human malarial parasites are also members of the Plasmodium genus.
“This work helps us to understand the transmission of these parasites between species and how that may play an important role in their evolution,” Weckstein explained. “Thus, it helps us to characterize the ecology of pathogen dispersal and host switching events, which is critical to understanding many of our own human pathogens.”
Drexel News is produced by
University Marketing and Communications.