Of all the drugs used in a hospital to treat patients, about 75% are secondary metabolite or secondary metabolite derived molecules. My lab is interested in all aspects of these metabolites, ranging from their biosynthesis and their microbiological role in nature to applications in the clinic.
Secondary Metabolism
Nature produces a wealth of compounds in primary and secondary metabolism. Traditionally, natural products, a key resource from secondary metabolism, have been studied at a chemical level before the gene clusters or enzymes responsible for their biosynthesis were known. Of late there is a strong trend in the opposite direction (“from gene to secondary metabolites”), especially strengthened by the recent technological breakthroughs in genome sequencing and bioinformatics. In my research group, we harvest a compilation of genomic, transcriptomic and metabolomic methods to explore fundamental questions within several lines of research.
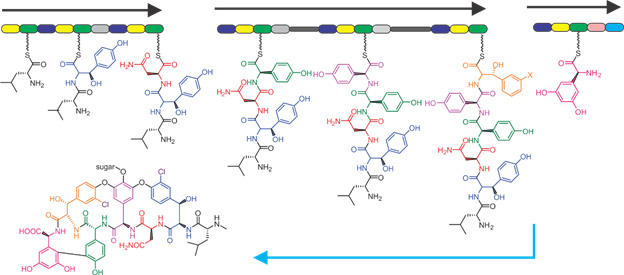
Biosynthetic gene cluster (BGC) of vancomycin, highlighting the conveyor-belt assembly line of this non-ribosomal peptide synthase producing the important antibiotic vancomycin, produced in Amycolatopsis orientalis.
Mass Spectrometry
Mass spectrometry, specifically tandem mass spectrometry (MS/MS) is the method of choice to mine complex samples for new natural products. Coupled with sensitive liquid chromatography we have a pipeline from genome to secondary metabolite. Gas chromatography MS is ideal for screening engineered bacterial strains for small metabolites like terpenes, fatty acids or small polyketides.

Liquid chromatography mass spectrometry setups in the New College Building include (from left to right) a Waters Acquity UPLC-Synapt G2Si QTOF, Thermo Eksigent 2d-nano-lc-LTQ linear ion trap, Waters Alliance HPLC-QDA single quad and Sciex Perkin Elmer-triple-quad (Waters Alliance-ZQD single quad not shown).
Pathogenic Bacteria
Pathogenic bacteria have a fascinating arsenal of virulence factors. We explore the properties of these factors in biological assays.
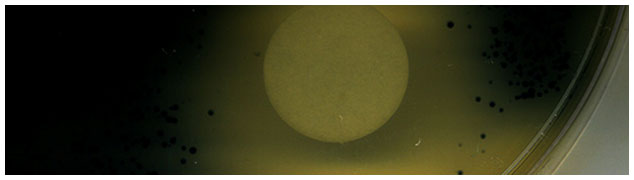
Bacterial colonies on an agar surface expressing a synthase that produces a deep blue dye. The filter disk (white circle) is spotted with a molecule from another bacterium and a zone of inhibition is observed.
Others
Many other exciting projects in the field of secondary metabolism, pathogenic bacteria and mass spectrometry are being pursued.